Latest Headlines
- What the New CMS Reimbursement for Principal Illness Navigation Means for Oncology Nurses
- Psychological Distress Linked to Quality of Life and Overall Survival in Patients Receiving AlloHSCT
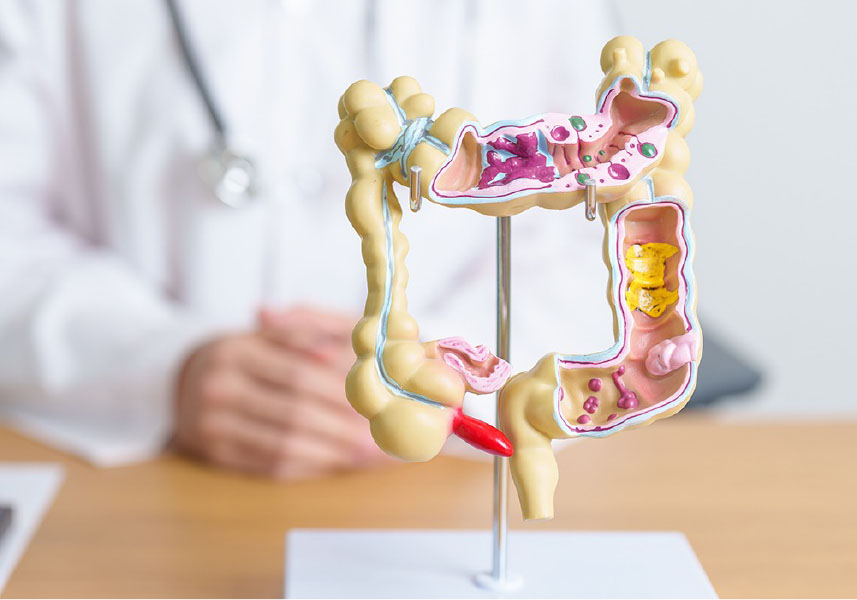
- Specialized Center Customizes Care for Growing Population of Patients With Young-Onset CRC
- FDA Grants Accelerated Approval to Fam-Trastuzumab Deruxtecan-Nxki for Unresectable or Metastatic HER2-Positive Solid Tumors
48th Annual ONS Congress® Newsreels
- Attendees Leave ONS Congress® and Prepare to Transform Cancer Care April 30, 2023
- Foundation Doubled Its Fundraising Totals at 48th Annual ONS Congress® April 30, 2023
- Thank You, ONS Congress® Attendees! April 30, 2023
- Viva Fiesta With the Local Chapter at ONS Congress® in San Antonio, TX April 29, 2023
- ONS Congress® Reception Celebrates Outgoing President Jeannine Brant April 29, 2023
Our Spirit. Our Practice.

My husband is a geriatrician.
My oldest daughter first became a music therapist and then completed an accelerated program in nursing. She is passionate about caring for older adults, just like her father. She completed her master’s degree in nursing and is now working as a palliative care nurse practitioner in a very busy urban university trauma center.